The frequent k-mers problems is one of the very first problems introduced to those who took Bioinformatics I course on Coursera. It is a problem which is not difficult, but also represents a good introduction to the field since its naive solution is impractical, and it requires a little bit of optimization. After solving the problem as efficiently as possible, I started thinking of implementing it as a MapReduce task for practice.
First, does the problem really need to be distributed? I do not know! My lack of experience does not allow me to know much about how frequently people need to find the most frequent k-mers. However, for the sake of practice, I decided to implement it,; there is no harm. In this post, we will not discuss how to solve the problem itself, but rather we will discuss how to properly split the input to be able to write the MapReduce task correctly.
Problem definition
In case you are not familiar with the problem, it is simply defined as: “given a string, find the most common sub-string of length k”
Sounds similar? the problem is very similar to the word count problem, but all words are of a fixed length, and the output is only the words which occurred most frequently.
Solution
The solution is divided into four parts: partitioning, mapping, reducing, and finding the final answer. In order to implement it, I will use LazyMapReduce, but you can achieve the same thing with any other MapReduce implementation.
Since we are already familiar with Map and Reduce for the word count problem, we can just write them directly. Notice that the two functions are independent of how partitioning will be done, as Map will only work on an individual partition. This time, we will make a class which extends MapReduceTask and assigns Map and Reduce internally, instead of creating a MapReduceTask instance and assigning its Map and Reduce functions.
Just like the word count count problem, the MapReduceTask types will be ‘<object, string, string, int, string, int>’ (refer to the previous tutorial to know more about that). Therefore the structure of the class will be like this
public class FrequentKmersTask: MapReduceTask<object, string, string, int, string, int>
{
string sequence;
int k;
int partitionSize;
public FrequentKmersTask (string _sequence, int _k, int _partitionSize)
{
sequence = _sequence;
k = _k;
partitionSize = _partitionSize;
this.Map = FrequentKmersMap;
this.Reduce = FrequentKmersReduce;
}
}
Now we will work on the very first part of the implementation, which is how to split the genome sequence into chunks, each to be pushed as input to the Map function.
Let’s just divide the genome into equal-sized partitions, where the last one might be shorter than the rest, and make every partition start right after the previous one.
void PrepareTaks ()
{
var index = 0;
var length = 0;
var partition = "";
while (true) {
length = Math.Min (partitionSize, sequence.Length - index);
partition = sequence.Substring (index, length);
PushInput (null, partition);
index += length;
if (index >= sequence.Length)
break;
}
}
That was easy, and we are done. NO, NOT SO FAST. There is a big flaw in the way we split the genome. A k-mer which is divided between two chunks will be ignored by our Map function. To make this clearer let’s try to check mathematically if the algorithm we use to partition a genome sequence will give us in the end the same number of k-mers or not.
First, we need to know how many k-mers a string of length N has, and it is  (you can try to verify that if you would like to). Then we split the string into m chunks with size n’, and according to our partitioning algorithm,
(you can try to verify that if you would like to). Then we split the string into m chunks with size n’, and according to our partitioning algorithm, 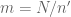 .
.
Then we need to calculate how many k-mers will result from splitting the string the way we did. The total number of k-mers is 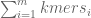 =
=  . Then after replacing
. Then after replacing 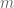 with $N/n’$ the formula can be rewritten as
with $N/n’$ the formula can be rewritten as 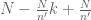 .
.
The only situation where  is when
is when 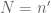 , which means that we do not split the genome at all. On the other hand, the solution is rather simple, and it only involves modifying one statement.
, which means that we do not split the genome at all. On the other hand, the solution is rather simple, and it only involves modifying one statement.
But before we go into the code, let’s fix the way we thought about splitting the string first, and change it to “divide the genome into equal-sized partitions, where the last one might be shorter than the rest, and make every partition start right after the LAST INDEX WHICH IS A VALID START OF A K-MER IN THE previous one”.
In a string of size 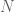 , the indices which are starts of k-mers are
, the indices which are starts of k-mers are 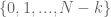 . Therefore, for chunks of size
. Therefore, for chunks of size  , each chunk will start right at
, each chunk will start right at  , where
, where 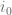 is the starting index of the previous chunk.
is the starting index of the previous chunk.
To achieve that we only need to change the way we update the starting index of the next chunk from the previous implementation, and convert it to
void PrepareTaks ()
{
var index = 0;
var length = 0;
var partition = "";
while (true) {
length = Math.Min (partitionSize, sequence.Length - index);
partition = sequence.Substring (index, length);
PushInput (null, partition);
index += partitionSize - k + 1;
if (index >= sequence.Length)
break;
}
}
And then, FrequentKmersMap and FrequentKmersReduce will be defined respectively as
List<KeySingleValuePair<string, int>> FrequentKmersMap (KeySingleValuePair<object, string> input)
{
string kmer;
var partition = input.value;
var result = new List<KeySingleValuePair<string, int>> ();
for (int i = 0; i <= partition.Length - k; i++) {
kmer = partition.Substring (i, k);
result.Add (new KeySingleValuePair<string, int> {
key = kmer,
value = 1
});
}
return result;
}
KeySingleValuePair<string, int> FrequentKmersReduce (KeyMultipleValuesPair<string, int> input)
{
return new KeySingleValuePair<string, int> {
key = input.key,
value = input.values.Count
};
}
Since everything is ready, the only remaining task is to find the most frequent k-mers. The results of all Reduce calls can be retrieved by calling GetResults(), then by iterating through them we can find the most frequent k-mers as follows
public List<string> GetFrequentKmers ()
{
var results = GetResults ();
var kmers = new List<string> ();
var maxCount = 0;
foreach (var kmer in results) {
if (kmer.value > maxCount) {
maxCount = kmer.value;
kmers.Clear ();
kmers.Add (kmer.key);
} else if (kmer.value == maxCount) {
kmers.Add (kmer.key);
}
}
return kmers;
}
considerably worse than
as an algorithm complexity? Should we strive to find the constant-time solution? There’s no definitive answer for that, and it usually is “depends on what you’re trying to achieve”. This is why I’m going to present the comparison in a more abstract sense, by looking at them.
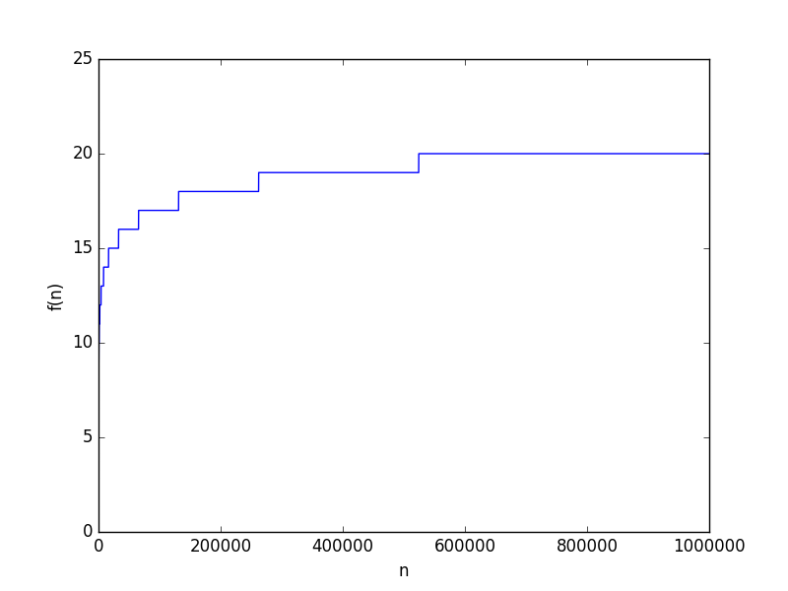
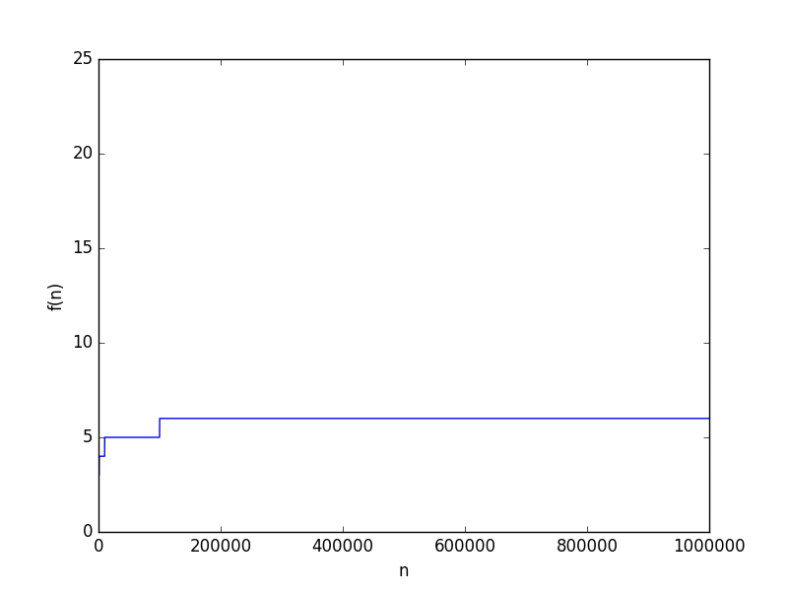
looks like
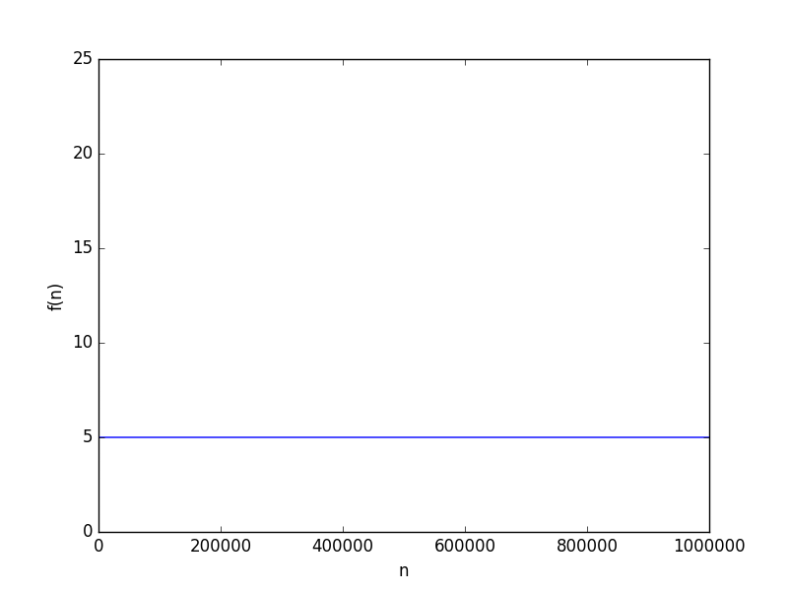
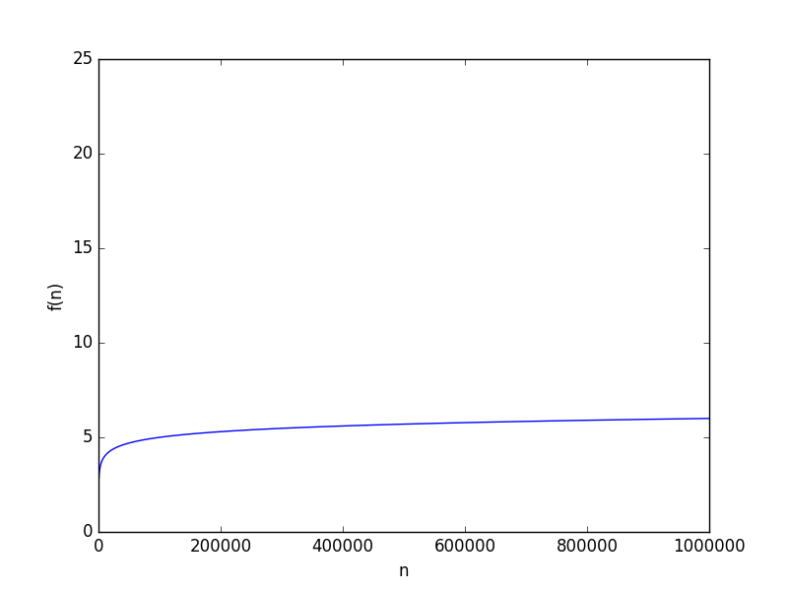
to hit 6. But this because the base was 10, how about a smaller base, say 2, it will be way worse, right? From a complexity point of view, the base of a logarithmic function is just a constant, and it just doesn’t matter when it comes for asymptotic growth (you can have some fun finding why it doesn’t).
to test anyway. It’s worse but not “way” worse at all. At
the value was just 20.
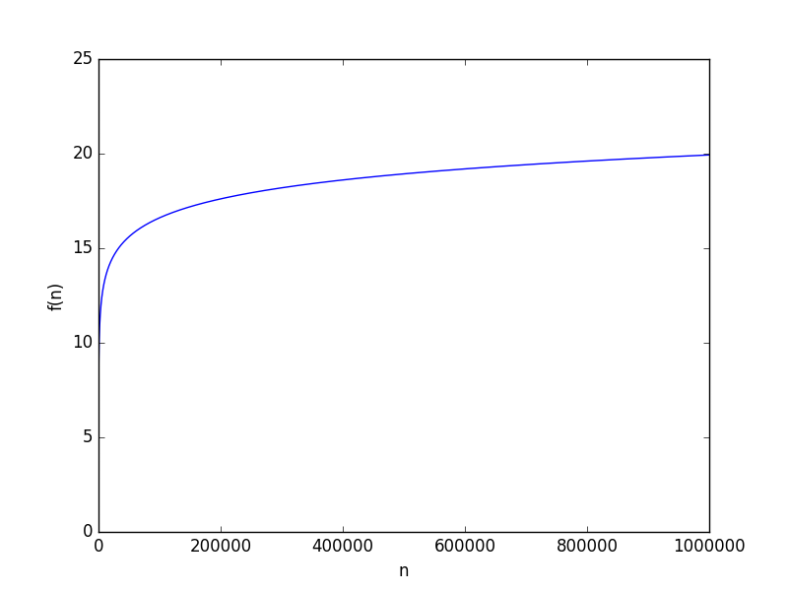
and
instead and see how close to constant they look yourself.